Biomedical journals are a rich source of information for such multimodal content indexing. Recent efforts in biomedical visual question answering (VQA) research rely on combined information gathered from the image content and surrounding text supporting the figure. Developing an interactive visualization tool that can display modeling results from a large materials network perspective as well as a time-based perspective is an advancement in visualization studies. Also, the field of image-based document analysis will benefit tremendously from machine learning tools such as the use of deep belief networks for classification and text separation from document images. The research will advance the machine-learning area of developing hierarchical, dynamic topic models to investigate trends in materials discovery over user-specified time periods. While the proposed methodology targets the interdisciplinary field of materials research, the tools to be developed can be generalized to enhance scientific discoveries and learning across a broad swathe of disciplines. So, the major thrust of this concept paper is the use of technology to (i) extract "deep" meaning from a large corpus of relevant materials science documents (ii) navigate, cluster, and present documents in a meaningful way and (iii) evaluate and revise the materials-related query responses until the researchers are guided to their information destination. To obtain insights for the discovery of new materials and to study about existing materials, research and development scientists and engineers rely heavily on an ever-growing number of materials research publications, mostly available online, and that date back many decades. This chapter presents a concept paper that describes methods to accelerate new materials discovery and optimization, by enabling faster recognition and use of important theoretical, computational, and experimental information aggregated from peer-reviewed and published materials-related scientific documents online. We present the successes and identify critical challenges. 
Experiments on 515 multi-panel figures and analysis of the results show promising results. We propose a 4-step panel label detection method based on Markov Random Field (MRF). Subfigure labels are valuable in associating individual panels with relevant text in captions and discussion. 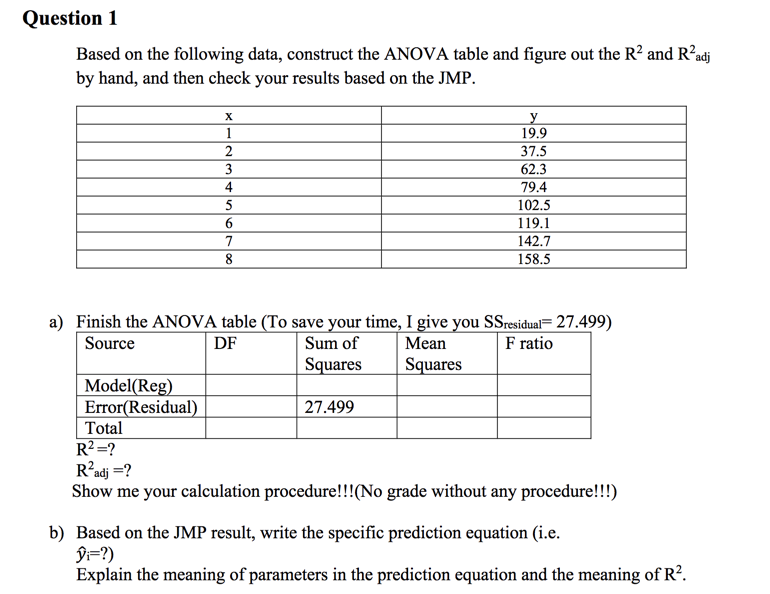
Prior to feature extraction for indexing and retrieval of biomedical figures it is necessary to classify image content in each subfigure by its modality (X-ray, MRI, CT, etc.) and other relevant criteria. Splitting such multi-panel figures into individual subfigures is a necessary step for improved multimodal biomedical information retrieval. ) which are referenced in the figure caption and discussion in the article body. Figures in biomedical articles often comprise several subfigures that are identified by superimposed panel labels ('A', 'B'. We present a method for figure-panel (subfigure) label detection and recognition in multi-panel figures extracted from biomedical articles. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
